In agricultural production, trait-related gene mapping is very important for scientific breeding. Traditional genetic recombination positioning has guiding significance and practical value, but traditional methods have long cycle and high cost. With the development of second-generation sequencing, DNA sequencing speed is greatly accelerated and the cost is greatly reduced, making the mapping-by-sequencing by sequencing method become the main means of more and more animal and plant research. For the appropriate sample population, combined with traditional cluster separation analysis (BSA) and second-generation sequencing methods, the trait-related genes can be quickly located under the complete genome reference sequence. In just a few days, the region of the gene can be defined by data manipulation to further lock the gene. This method has been applied to the genetic mapping of Arabidopsis, rice, maize, mice, zebrafish, and the like.
Usually the premise of this application requires a high quality, complete reference genomic sequence, including all genes and other genomic sequence information. However, the genomic maps of many important crops in agriculture, such as barley, wheat, etc., are not complete. German scientists recently published an article in Genome Biology 1, introduced by exome sequencing method for positioning a second generation effects of barley leaf out rate (leaf initiation rate) gene, can also be shown that even in the case of incomplete genome Gene mapping for sequencing.
This study is aimed at the barley multi-segment dwarf mutant (mnd) . The most prominent feature of this mutant is the shortening of the inter-leaf period, the speed of leaf emergence , the average number of leaves is twice that of the wild type, and the length of the internode is shortened. And accompanied by changes in the shape of the leaves and so on. The F2 generation was obtained by X-ray-induced multi-segment dwarf mutant hybridization with wild-type cv. Barke. In 100 F2, 19% were mutant and 81% were wild type, and the trait was a single gene recessive inheritance. The DNA of 18 mutants was mixed into a DNA mixture of a mutant sample population, and then a random mixture of 30 wild-type mixed wild-type sample populations was used. The NimbleGen barley exome probe was used for the two DNA mixtures. After enrichment of the coding sequence of the barley genome, second generation sequencing was performed.
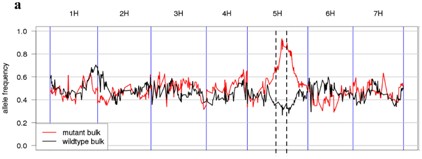
Figure: Gene mapping: Candidate genes were mapped by differences in allele frequencies between the mutant sample population and the wild sample population. From Mascher M, Jost M, Kuon JE, Himmelbach A, Aßfalg A, Beier S, Scholz U, Graner A, Stein N. Mapping-by-sequencing accelerates forward genetics in barley. Genome Biol. 2014. Jun 10;15(6 ):R78
The mutated sample population and the wild-type sample population were obtained with 8.2 Gb and 7Gb sequencing data, respectively, and the SNP locus and its allele frequency were analyzed after cv. Barke's genome-wide shotgun sequencing. Comparing the SNP locus allele frequencies of the two sample groups, a 30 cM region was found in the long arm of chromosome 5H. The frequency of the mutant SNP allele was 95% in this region, while the wild-type SNP frequency was only 30%. At the same time, since the mutations caused by X-rays are mostly deletion mutations, a deletion mutation of >150 bp was found by sequencing in the sequencing data, and 18 deletions were found in the above region. In the contig corresponding to the deletion fragment, contig 49382 is located at 5H 96 cM and has two deletion regions, which are two exons of the MLOC_64838.2 gene annotated as "Cytochrome P450". Blastn's results showed that the protein encoded by this gene is similar to the protein sequence encoded by the rice PLASTOCHRON1 ( pla1 ) gene. The mutation of the pla1 gene will make the rice have a phenotype similar to that of the barley multi-segment dwarf mutant. The authors believe that the MLOC_64838.2 gene is likely It is a gene that controls this shape. The article also validated this gene in other barley multi-segment dwarf mutants and mapped the gene in the physical map of barley. The authors believe that the use of a large number of existing barley mutant resources, combined with exome sequencing, rational data analysis methods and existing genomic resources, can cost-effectively map the sequencing of barley by sequencing. The method can also be applied to similar studies of large genomes of other genomes.
The NimbleGen barley exome probe design used in this paper is based on the annotation of the existing barley genome splicing, and is designed with reference to the full-length RNA sequence and the RNASeq sequencing sequence, for a total of about 88.6 Mb target sequence, corresponding to ~60 Mb in the current barley genome. Encoding area. The barley exome capture product has approximately 4.2 million different DNA probes in a capture reaction solution, demonstrating the NimbleGen high-density probe production and design capabilities to effectively cover and capture target areas. A paper published in The Plant Journal in November 20132 also used the barley exome capture product to conduct experiments on different strains of barley and wild barley, followed by SNP discovery, diversity analysis and evolution analysis. The article pointed out that this tool can be applied to barley and even wheat of various strains, which helps the variation of mRNA coding region, greatly reduces the research cost by mixing capture and reducing the amount of sequencing, and thus promotes the study of barley genetic diversity and genetic traits.
1. Mascher M, Jost M, Kuon JE, Himmelbach A, Aßfalg A, Beier S, Scholz U, Graner A, Stein N. Mapping-by-sequencing accelerates forward genetics in barley. Genome Biol. 2014. Jun 10;15( 6): R78.
2. Mascher M, Richmond TA, Gerhardt DJ, Himmelbach A, Clissold L, Sampath D, Ayling S, Steuernagel B, Pfeifer M, D'Ascenzo M, Akhunov ED, Hedley PE, Gonzales AM, Morrell PL, Kilian B, Blattner FR, Scholz U, Mayer KF, Flavell AJ, Muehlbauer GJ, Waugh R, Jeddeloh JA, Stein N. Barley whole exome capture: a tool for genomic research in the genus Hordeum and beyond. Plant J. 2013 Nov;76(3) :494-505. doi: 10.1111/tpj.12294. Epub 2013 Aug 24.
For scientific research only, not for clinical diagnosis
Fresh Half Shell Mussel Meat,Half Shell Mussel Meat,Frozen Cooked Mussel Meat,Frozen Mussel
Shengsi Huali Aquatic Products Co.,Ltd , https://www.mytilus-edulis.com