Nie Aiying
Thermo Fisher Scientific (China) Co., Ltd.
Key words
Q-Exactive; ADC monoclonal antibody; molecular weight determination; amino acid sequence analysis; glycosylation site analysis; ADC drug binding site analysis
1 Introduction
Monoclonal antibody drugs are highly specific targeted drugs that specifically act on tumor cells and are known as "bio-guides" for the treatment of malignant tumors. The ADC antibody is a non-targeted chemotherapeutic drug (or small molecule drug) that is anti-tumor on a specific natural amino acid of the antibody protein to increase the efficacy of the monoclonal antibody and reduce the side reaction of the small molecule drug. In this regard, the researchers said that this is equivalent to accurately mounting the "nuclear warhead" on the "bio-missile", making treatment safer, more effective, and more oriented. At present, ADC antibody drugs have been widely recognized in the field of oncology research. Many multinational pharmaceutical companies have successively devoted themselves to the research and development of ADC drugs. In order to ensure the safety and efficacy of the ADC monoclonal antibody and the real-time monitoring of the monoclonal antibody production process, the determination of the overall molecular weight of the ADC monoclonal antibody, the identification of the amino acid sequence, the confirmation of the glycosylation site, and the ADC drug The testing and confirmation of important ADC drug analysis indicators such as confirmation of binding sites, localization of disulfide bonds, and analysis of sugar chain structure are increasingly attracting attention from major pharmaceutical companies. At present, with the continuous development and popularization of bio-mass spectrometry technology, the single-antibody analysis technology based on bio-mass spectrometry has begun to be comprehensively and rapidly established.
Q-Exactive tandem mass spectrometer quadrupole high selectivity and high resolution mass spectrometry, high sensitivity mass analyzer Orbitrap TM, greatly improving the selectivity, the sensitivity and accuracy of mass of the ions. With 140,000 resolution (FWHM), higher sensitivity, faster scanning speed and greater dynamic range, the system combines HCD fragmentation technology to enable cascade mass spectrometry fragmentation and identification for deep complex samples. Analysis, including proteomics, metabolomics, lipidomics and other samples, post-translational modification and quantitative analysis, provides a comprehensive, efficient and fast mass spectrometry platform. Similarly, Q-Exactive mass spectrometry can be effectively applied to the development and production monitoring of ADC monoclonal antibodies. The basic experimental analysis process is as follows.
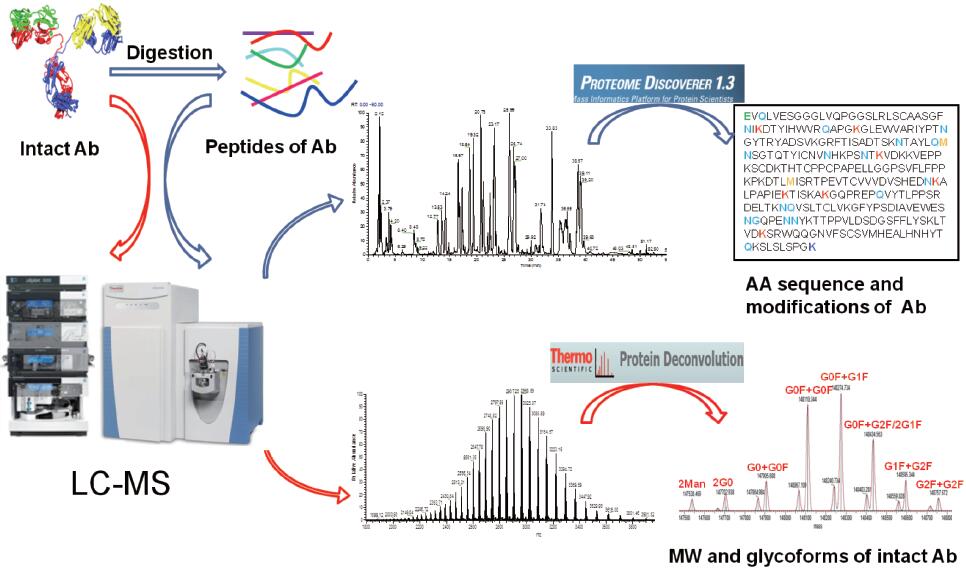
Figure 1. Basic flowchart of monoclonal antibody analysis
2. Experimental part
2.1 Determination of molecular weight
2.1.1 Instruments and reagents
Mass Spectrometry Instruments: Q-Exactive (Thermo Fisher Scientific, USA);
Chromatography instrument: Accela LC system (Thermo Fisher Scientific, USA);
Column: Agilent Zorbax 300, SB-C8, 3 μm, 2.1 × 50 mm;
Reagents: secondary deionized water, chromatographic grade acetonitrile, chromatographic grade formic acid.
2.1.2 Instrument method
Chromatographic conditions: see Table 1 for details;
Mass spectrometry conditions: see Table 2 for details;
Table 1. Chromatographic conditions for molecular weight determination of monoclonal antibodies
Mobile phase | A: 0.1% formic acid aqueous solution; B: 0.1% formic acid acetonitrile solution | |
Flow rate | 300 μL/min | |
Column temperature | 75 ° C | |
Chromatographic gradient | Time/min | Phase B concentration /% |
0 | 5 | |
4 | 5 | |
12 | 90 | |
12.1 | 90 | |
16 | 90 | |
16.5 | 5 | |
20 | 5 | |
Table 2. Mass spectrometric conditions for molecular weight determination of monoclonal antibodies
Spray voltage | 4 kV |
Capillary heating temperature | 275 ° C |
S-lens | 60%; |
Sheath gas flow rate | 30(arb) |
Auxiliary gas flow rate | 5(arb) |
Quality scan range | m/z 2000–4000 |
Resolution | 17,500 (m/z 200) |
Intra source lysis (SID) | 100 eV |
Microscan | 10 |
2.1.3 Data Analysis Method
The original mass spectrum was deconvoluted using Protein Deconvolution 2.0 software to obtain complete protein molecular weight information.
2.2 Amino acid sequence, glycosylation site and ADC drug binding site determination
2.2.1 Instruments and reagents
Mass Spectrometry Instruments: Q-Exactive (Thermo Fisher Scientific, USA);
Chromatography instrument: Accela LC system (Thermo Fisher Scientific, USA);
Column: Thermo Scientific, C18, 100Å, 1.9 μm, 2.1×100 mm
Reagents: secondary deionized water, chromatographic grade acetonitrile, chromatographic grade formic acid.
2.2.2 Instrument method
Chromatographic conditions: see Table 3 for details;
Table 3. Chromatographic conditions for amino acid sequence determination
Mobile phase | A: 0.1% formic acid aqueous solution; B: 0.1% formic acid acetonitrile solution | |
Flow rate | 300 μL/min | |
Column temperature | 35°C | |
Chromatographic gradient | Time/min | Phase B concentration /% |
0 | 5 | |
4 | 5 | |
45 | 35 | |
50 | 90 | |
52 | 90 | |
52.5 | 5 | |
60 | 5 | |
Mass spectrometry conditions: see Table 4 for details;
Table 4. Mass spectrometric conditions for amino acid sequence determination
Spray voltage | 3.5 kV |
Capillary heating temperature | 275 ° C |
S-lens | 60%; |
Collision energy | 27% normalized energy |
Fragmentation | HCD |
Quality scan range | m/z 300–1800 |
Resolution | Level 35,000 (m/z 200), |
Level 17,500 (m/z 200) |
2.2.3 Data analysis method
The original spectrum was searched using Proteome Discoverer 1.3 software. The specific search parameters were: database containing the monoclonal antibody amino acid sequence; cysteine ​​(C) alkylation (+57.021Da) was set as a fixed modification; methotrexate Acid (M) oxidation (+15.995 Da), asparagine (N) and glutamine (Q) deamination (+0.984 Da), lysine (K) ADC binding (+956.3644 Da), asparagine (N) G0F glycosylation (+1444.5300 Da) was set to variable modification; trypsin was set to enzyme; enzyme cleavage site was 2.
3. Results and discussion
3.1 Determination of protein molecular weight
Based on Q-Exactive, we set the Orbitrap detector's acquisition resolution to 17,500 to ensure the quality and quantity of the acquired spectra. The total chromatogram of the original chromatographic mass spectrum of the ADC monoclonal antibody collected by Q-Exactive mass spectrometry and the original mass spectrum are shown in Figures 2 and 3. We can observe that the ADC antibody drug exhibits a uniform charge distribution over the entire mass range. Baseline separation can be achieved between mass peaks of different charges, so that the sensitivity, resolution, and signal-to-noise ratio of the mass spectrum are high. At the same time, several sets of mass spectral peaks with higher intensity are selected for amplification, and the mass spectrum peaks after amplification show similar distribution patterns.
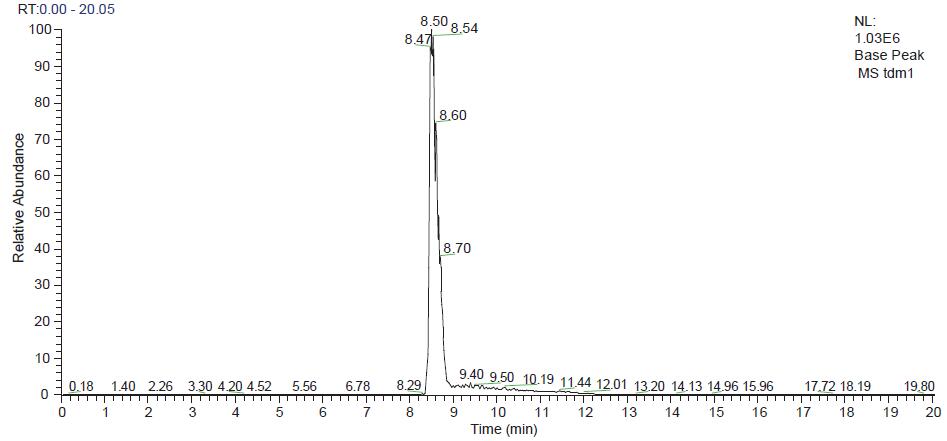
Figure 2. ADC antibody drug chromatographic mass spectrometry outflow map
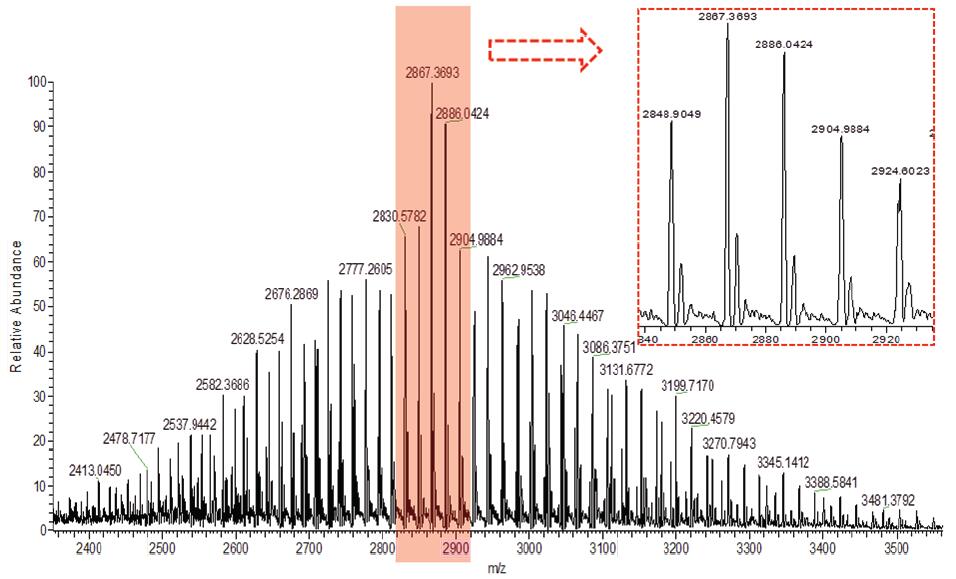
Figure 3. Original spectrum of ADC antibody drug (magnified view of the middle set of peaks in the red dashed box)
The molecular weight distribution of the monoclonal antibody after deconvolution by Protein Deconvolution 2.0 software is shown below (Fig. 4), and the observed mass spectrum peaks are calculated according to the molecular weight of the amino acid theoretical sequence of the monoclonal antibody, and we observe from the figure There is a mass increase of about 956 Da between the ADC monoclonal antibodies, and it is concluded that the monoclonal antibody binds to different numbers of small drug molecules with a molecular weight of about 956 Da, and can detect up to 6 small drug molecules. . At the same time, an increase in mass of around 160 Da was observed, mainly due to the presence of the linking group. Therefore, it is preliminarily concluded that the ADC monoclonal antibody mainly combines six ADC small molecule drugs, and the result is basically consistent with the theoretical value. According to the peak intensity information of the mass spectrum peak, we can also calculate the ADR value of the drug, which is for the ADC. Evaluation of the effectiveness of a drug is critical.
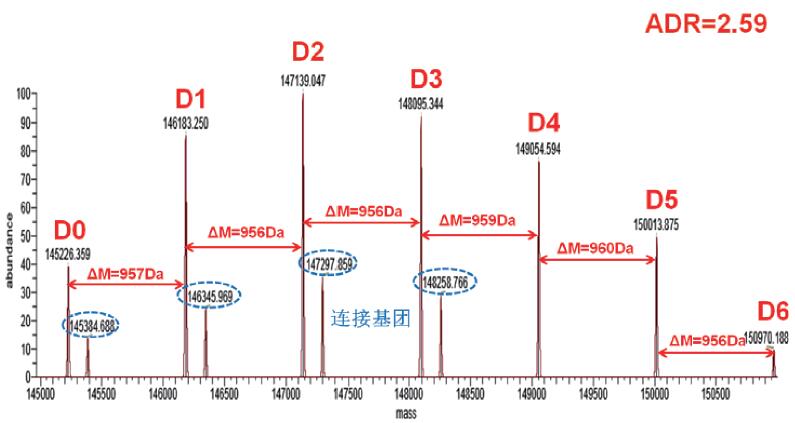
Figure 4. The original mass spectrum of the ADC mAb captured by Q-Exactive mass spectrometry.
Molecular Weight Distribution of Monoclonal Antibody after Deconvolution 2.0 Software Deconvolution
(D0–D6 represents a variant of a monoclonal antibody that binds to a small molecule of a different ADC drug, and a blue circle represents the mass spectral peak corresponding to the linking group.)
3.2 Amino acid sequence, glycosylation site and ADC drug binding site identification
3.2.1 Amino acid sequence coverage
The chromatographic mass spectrometry elution pattern obtained by trypsin digestion on Q-Exactive mass spectrometer is shown in Figure 5. After searching through the SEQUEEST database, the protein sequence coverage of the ADC monoclonal antibody heavy chain is 90.24%, such as As shown in Fig. 6(A), the protein sequence coverage of the light chain is 100%, as shown in Fig. 6(B), wherein the unidentified amino acid sequence is mainly due to the lack of an enzyme cleavage site and the enzymatic cleavage of the peptide segment. In short, in order to achieve 100% amino acid sequence coverage, it can be achieved by selecting other endoproteases.
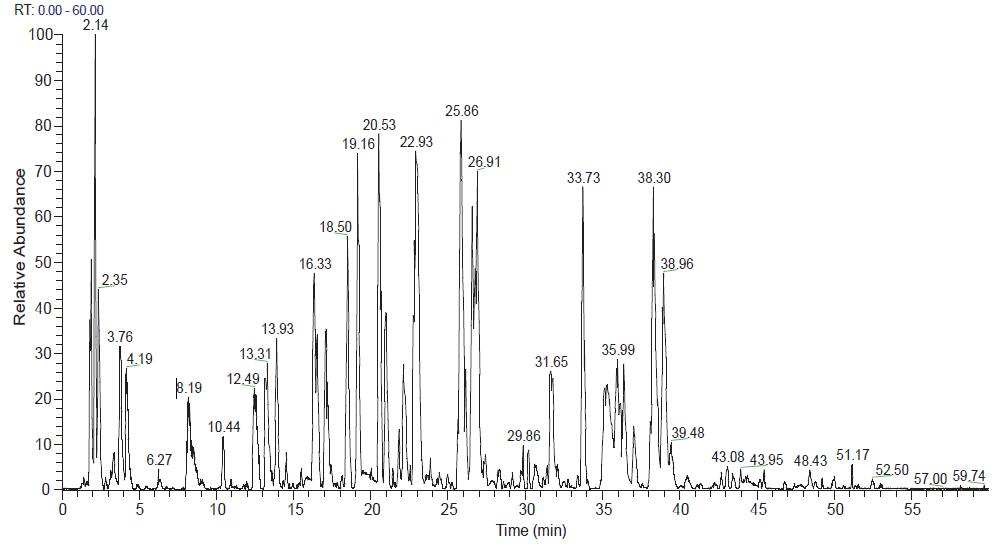
Figure 5. Chromatographic mass spectrometry outflow of the Trypsin enzymatic peptide of the ADC monoclonal antibody
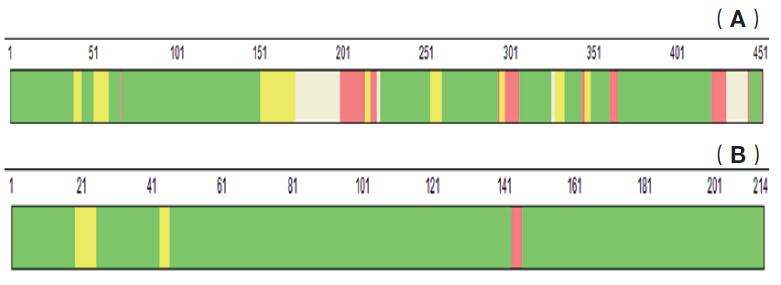
Figure 6. Schematic diagram of amino acid sequence coverage of ADC antibody drugs by trypsin digestion and Q-Exactive mass spectrometry. (A is a schematic diagram of the heavy chain amino acid sequence coverage, B is a light chain amino acid sequence coverage diagram, wherein the green labeled amino acid sequence indicates that the reliability of the sequence identification is very high, reaching more than 99%, and the red labeled amino acid sequence indicates the sequence The reliability of the identification is below 95%. The color-free amino acid sequence indicates that the sequence has not been identified, and the number above the amino acid sequence indicates the position of the amino acid.)
3.2 Determination of glycosylation sites
The ADC monoclonal antibody is identical to other monoclonal antibodies, and also contains N-glycosylation modifications, generally occurring in the conserved sequence NXS or NXT (X is any amino acid except proline). In the case of intact sugar chain, trypsin enzymatic hydrolysis is carried out directly. We can perform the G0F glycosylation variable modification setting during the search, and the N glycosylation site can be directly obtained: N261 in the heavy chain amino acid sequence EEQY N261STYR Site, the lower panel shows the G0F cascade mass spectrum of the peptide. Except for the b and y fragment ions of the peptide represented by yellow and blue in the figure, the red circle shows the sugar characteristic fragment ion of the glycopeptide. (low m/z end) and glycopeptide fragment ions (high m/z end), so that the presence of the glycopeptide and glycosylation site can be accurately confirmed.
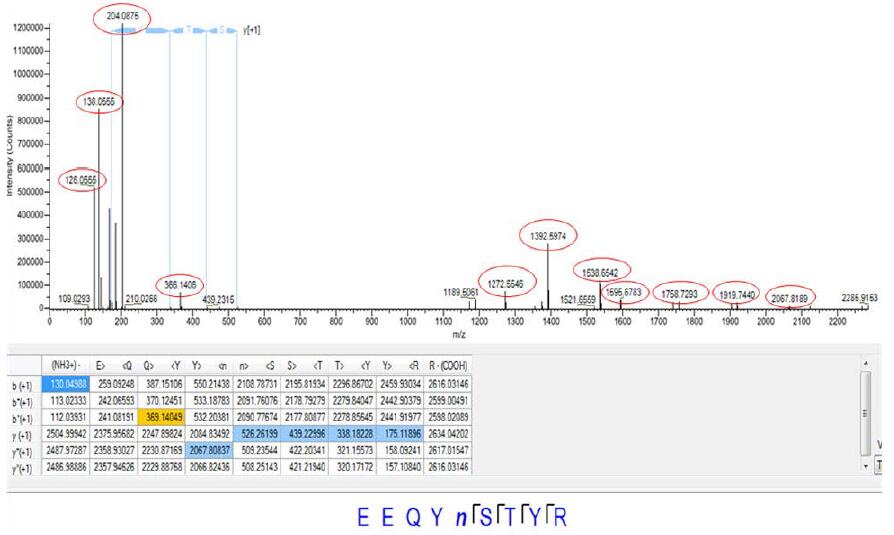
Figure 7. Cascade mass spectrum of the heavy chain amino acid sequence EEQYN261STYR
3.3 ADC drug binding site determination
According to the results of the Proteome Discovery software search, we found that there are characteristic fragment ions 453.20, 485.23 and 547.21 in the tandem mass spectrum of the peptides of lysine-binding drug molecules, so we can couple all possible ADCs through these feature fragments. The tandem mass spectrum of the peptides was manually confirmed, and finally it was confirmed that two lysine sites in the light chain and seven lysine sites in the heavy chain were combined with ADC small molecule drugs. The specific sites are as follows. (Figure 8), and taking one of the peptides as an example, gives a tandem mass spectrum of the ADC-binding peptide (Figure 9), and the red circle represents the ADC characteristic fragment ion. At the same time, we also observed that the hydrophobicity of the peptides after ADC drug binding is relatively strong, and the chromatographic efflux is relatively backward. This phenomenon is completely in line with theoretical speculation.

Figure 8. Schematic diagram of ADC drug small molecule binding sites
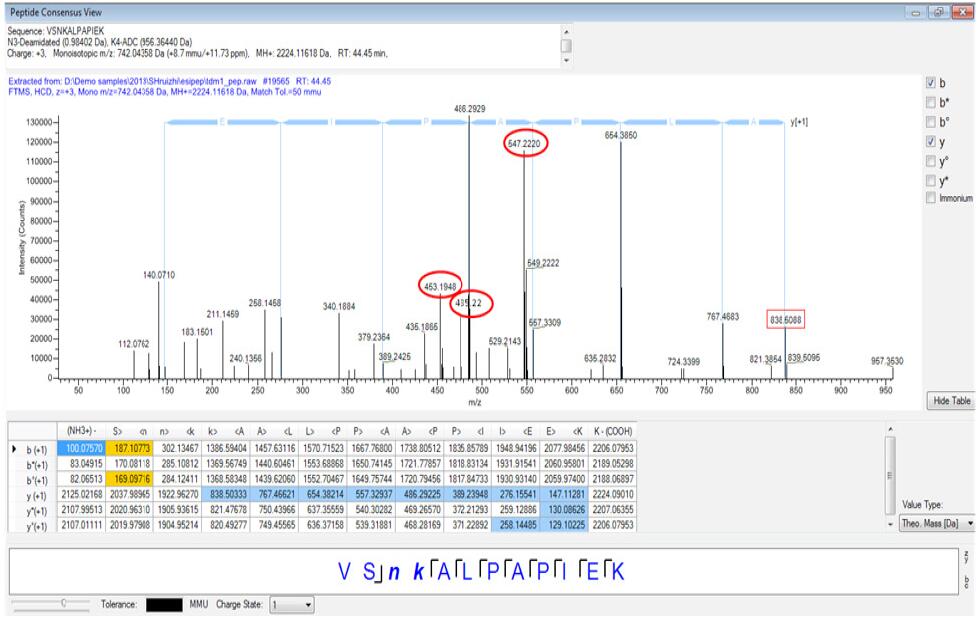
Figure 9. Cascade mass spectrum of the ADC binding peptide VSNKALPAPIEK
4 Conclusion
In this paper, Q-Exactive mass spectrometry was used to establish an analysis method for the overall molecular weight determination, amino acid sequence identification, glycosylation site and ADC drug binding site confirmation of ADC monoclonal antibody, which provided an analysis and real-time production detection for ADC monoclonal antibody. An efficient and fast analysis platform. The experimental results show that the Q-Exactive tandem mass spectrometer greatly improves and promotes the ADC monoclonal antibody with its ultra-high resolution, ultra-fast scanning speed, high mass accuracy, ultra-low sensitivity and large dynamic range. Identification analysis.


Orbitrap Group Club Thermo Fishery Mass Spectrometry Application Technology Group
Sweetness Polydextrose,Functional Food Ingredient,Low Sweetness Polydextrose
Shandong Bailong Chuangyuan Bio-tech Co.,Ltd. Qingdao Branch , https://www.sdblcycn.com